Bioscience
Short regulatory gene spotted
Two proteins produced by a single gene interact to keep the genome in check.
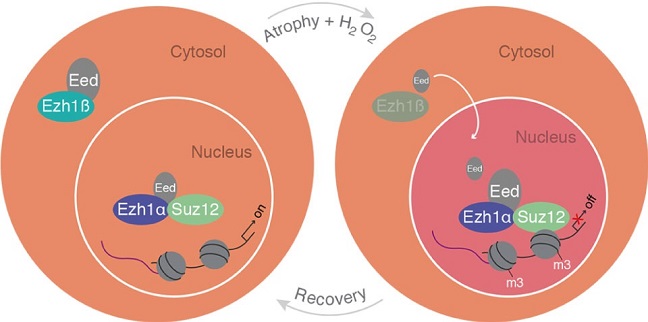
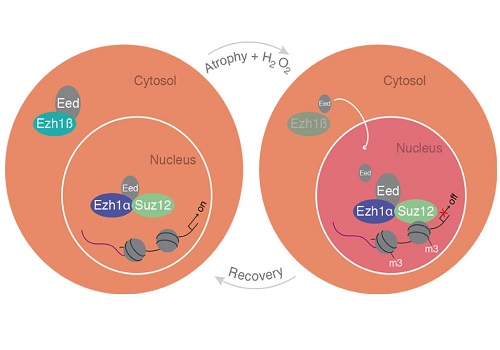
A schematic model of cytosolic Ezh1ß function in postmitotic differentiated muscle cells.
© KAUST 2017 Ivan Gromicho
An epigenetic mechanism regulating gene activity has been revealed by a KAUST-led international team of researchers investigating interactions between the human genome and its environment in adult tissues1.
Valerio Orlando’s lab at KAUST looks at the role of Ezh1, a gene whose function in mature tissues has remained unclear for 25 years. Like its twin Ezh2, Ezh1, along with a partner protein, encodes a protein involved in tagging genes to repress their activity. However, while Ezh2 mutations have been linked to cancer and developmental defects, mice lacking Ezh1 seem to develop normally.
Several years ago, Orlando’s group observed Ezh1 attached to the promoter of many genes that are normally switched on. “We saw this prototypical epigenetic repressor sitting on active genes, and our interpretation was that it’s there to provide the ability to repress them,” said Orlando. Hypothesizing that repression might be useful under stress, the team chemically stressed muscle cells and observed repression only in cells expressing Ezh1. Stress spurred Ezh1 into action, tagging genes with a repressive marker that could later be removed, a reversible response that Orlando calls “cell plasticity”: the ability to adapt to a dynamic environment.
A turning point in the conception of Ezh1 came when the team discovered a truncated version of the protein. Many human genes encode several slightly different versions of a protein, known as isoforms, and the researchers realized that an additional band lurking in some images was in fact a shorter isoform of Ezh1.
“Once our eyes were redirected to the short version, we immediately understood a number of things,” recalled Orlando. The truncated isoform was in the cytoplasm rather than the nucleus, and the team demonstrated that it acts as an environmental sensor regulating the activity of the full-length protein. Ezh1 needs a partner protein in order to tag genes, but the short isoform binds to the partner, trapping it in the cytoplasm, “like keeping that protein on a leash.” In stressed cells, the short isoform is degraded, releasing the partner to join full-length Ezh1 in the nucleus. Once the stress stops, short-Ezh1 once again traps the partner, stopping long-Ezh1 from acting, and the repressive tags are removed.
These findings reveal a new landscape of genetic regulation for researchers to explore, where interactions occur between isoforms of a single gene rather than products of different genes. “This offers a new paradigm for gene regulation, linking the genome with the environment,” said Orlando. “It’s a very exciting perspective.”
References
-
Bodega, B., Marasca, F., Ranzani, V., Cherubini, A., Della Valle, F … & Orlando, V. A cytosolic Ezh1 isoform modulates a PRC2-Ezh1 epigenetic adaptive response in postmitotic cells. Nature Structure & Molecular Biology 24, 444–452 (2017).| article
You might also like

Bioengineering
Bio-inspired network structures for next-generation AI

Bioscience
Robust workflow built for chemical genomic screening

Bioscience
Cell atlas offers clues to how childhood leukemia takes hold

Bioscience
Hidden flexibility in plant communication revealed

Bioscience
Harnessing the unintended epigenetic side effects of genome editing

Bioscience
Mica enables simpler, sharper, and deeper single-particle tracking

Bioengineering
Cancer’s hidden sugar code opens diagnostic opportunities
Bioscience



