Statistics
Stimulating deeper insights into brain function
Modeling changes in brain activity over time provides deeper insights into learning and behavioral responses.
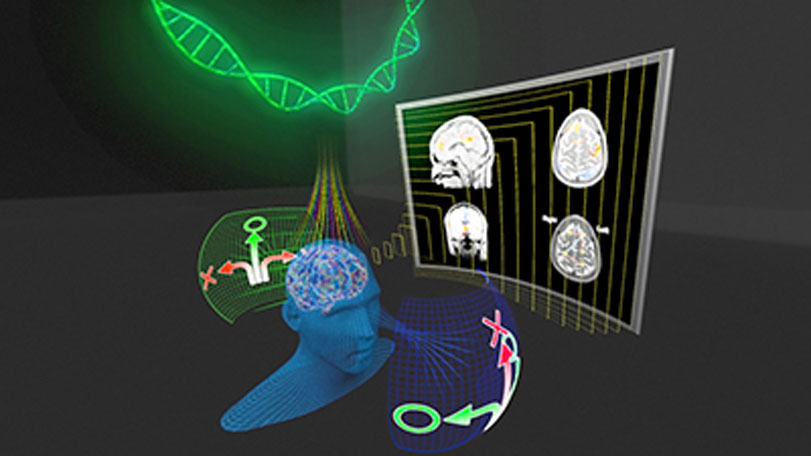
Observing the brain’s response to repeated stimuli has helped KAUST researchers develop a method for modeling connectivity patterns in neural networks. Mapping connectivity patterns will help to better understand brain function, ultimately improving diagnosis and treatment of brain diseases and mental disorders.
Neuroscientists believe that changes in brain activation and connectivity are associated with learning and habituation—the brain’s response to repeated stimulus evolves over time. However, there are currently no statistical models that can rigorously characterize this response.
Now, Hernando Ombao from KAUST, in collaboration with Mark Fiecas from the University of Minnesota in the United States and Chee-Ming Ting from the Universiti Teknologi Malaysia, has developed a robust statistical method for characterizing this evolving process.
“Our aim was to uncover dynamic changes in brain networks where interactions between brain regions exhibit changes over time,” says Ombao. “Characterizing the organization of brain networks provides fundamental insights into human behavior, cognition and emotion.”
Traditionally, neuroscientists have performed multiple trials to measure variations in connectivity patterns across the brain in response to repeated stimuli; they then averaged these results across trials. However, this approach doesn’t account for variations across trials due to systematic changes that may arise from habituation or changes that accompany learning.
To overcome this limitation, the researchers developed two general approaches for modeling dynamic brain connectivity that use imaging data from electroencephalograms—electrodes are placed on the scalp to record the brain’s electrical activity.
The first approach, called the Markov regime-switching vector autoregressive model, identifies a sequence of distinct, repeating connectivity patterns that are driven by a finite number of evolving brain states. While the second, known as the slowly evolutionary locally stationary process, captures the dynamic nature of the amplitudes of oscillatory activities of the brain at different frequencies.
“Our approach takes into consideration any subtle changes that may occur over the trials and could provide a better understanding of the ongoing cognitive processes over multiple timescales,” says Ombao.
While this work has successfully identified two potential approaches, it may also assist with development of more sophisticated and dynamic statistical models that better capture the structure of brain networks.
The researchers anticipate the application of this research in brain health. “Understanding brain connectivity patterns could be potentially useful as biomarker for the diagnosis of brain diseases and mental disorders, such as Alzheimer’s disease, obsessive compulsive disorder and schizophrenia, as well as leading to better treatments,” says Ombao.
References
- Ombao, H. Fiecas, M., Ting, C-M. & Low, Y.F. Statistical models for brain signals with properties that evolve across trials. NeuroImage 18, 609-618 (2018).| article
- Project Source Code https://github.com/marcoapintoo/kaust.kmindconnect
You might also like

Statistics
Checking your assumptions
Statistics
Internet searches offer early warnings of disease outbreaks
Statistics
Joining the dots for better health surveillance

Statistics
Easing the generation and storage of climate data

Statistics
A high-resolution boost for global climate modeling

Applied Mathematics and Computational Sciences
Finer forecasting to improve public health planning

Bioengineering
Shuffling the deck for privacy
Bioengineering




