Environmental Science and Engineering
Bringing together the tools to harness the potential of microbial diversity
Combining approaches to better discover and cultivate microbial taxa could generate novel microbes for industrial and agricultural applications.
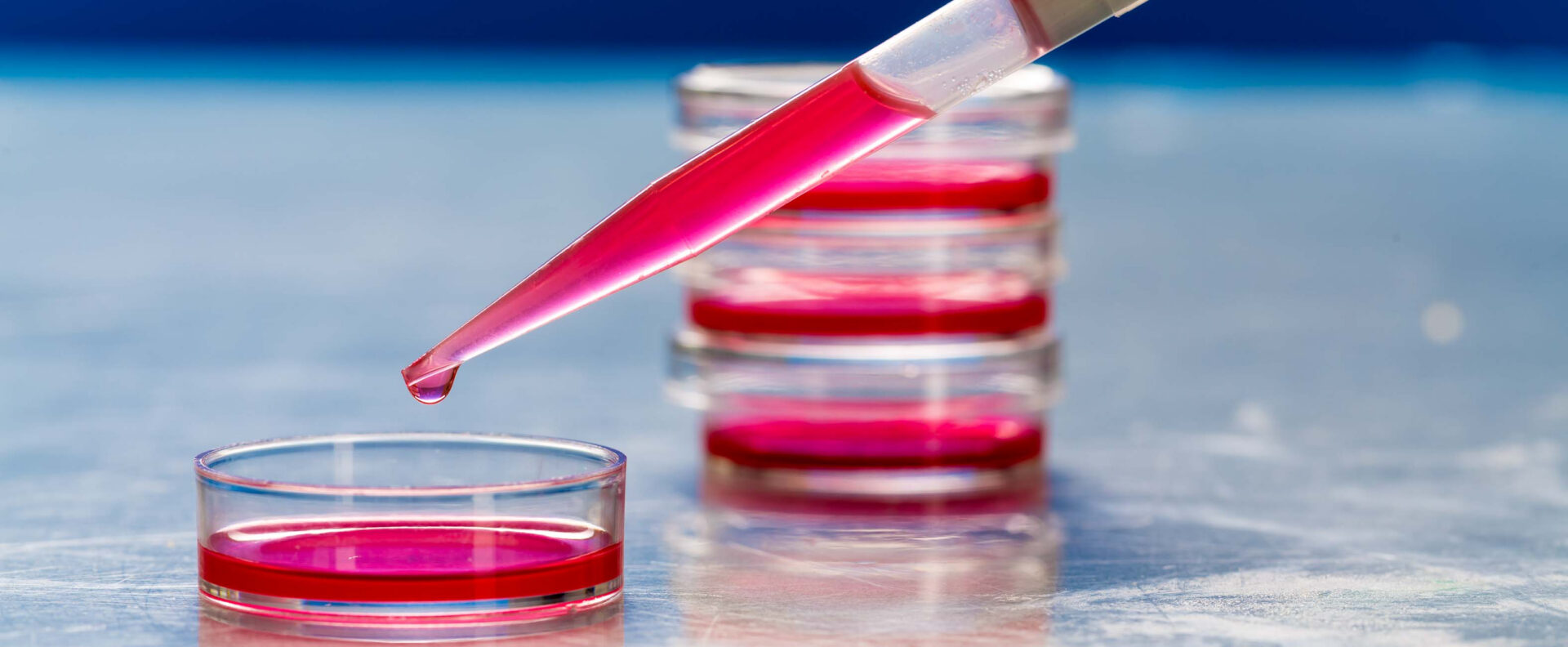

Advances in DNA sequencing have expanded our view of the microbial world, but the inability to cultivate most microbes has been a major constraint. Now, a systematic, predictive framework that combines existing genomic and computational modeling approaches to accelerate the discovery and cultivation of novel prokaryotic taxa has been proposed by KAUST researchers, in collaboration with an international team of scientists[1].
“Prokaryotic microbes — bacteria and archaea — are extraordinary microorganisms with unique capabilities that are already proving useful in industrial settings and biotechnological applications,” says Diego Javier Jiménez Avella, who led the perspective study within the research group of Alexandre Rosado at KAUST. “For example, cultivated microorganisms can be used in bioremediation processes, generating bioactive compounds, and enhancing plant growth in agricultural systems.”
Current DNA sequencing technologies have unveiled an enormous diversity of microbes on a global scale, but many potentially useful prokaryotic species remain undiscovered and uncultivated. Isolating and cultivating novel microbial species remains a major bottleneck in microbiology, notes Jiménez.
“We are at a very interesting time in microbiology, where we are revealing a vast microbial world through DNA, but we still cannot access much of it experimentally,” says Rosado. “Without cultivation, we cannot properly understand how these organisms function, or translate that knowledge into applications. There is also an urgency here: environments across the globe are changing rapidly, and we risk losing microbial diversity before we have had a chance to characterize it.”
Cultivation challenges
Scientists still do not fully understand the environmental and nutritional requirements needed to sustain microbial species in laboratory settings. Each microbe requires a unique combination of factors to grow, and this challenge is compounded by competition and interactions between microbial species.
“A wide range of techniques have been developed to isolate and cultivate microbial species from natural environments, but most of these rely on set nutrient and environmental conditions that tend to favor fast-growing, heterotrophic microorganisms,” says Jiménez. “Moreover, conventional isolation methods often disrupt obligate microbial interactions,” adds Jiménez, referring to the close interdependent relationships on which some species depend.
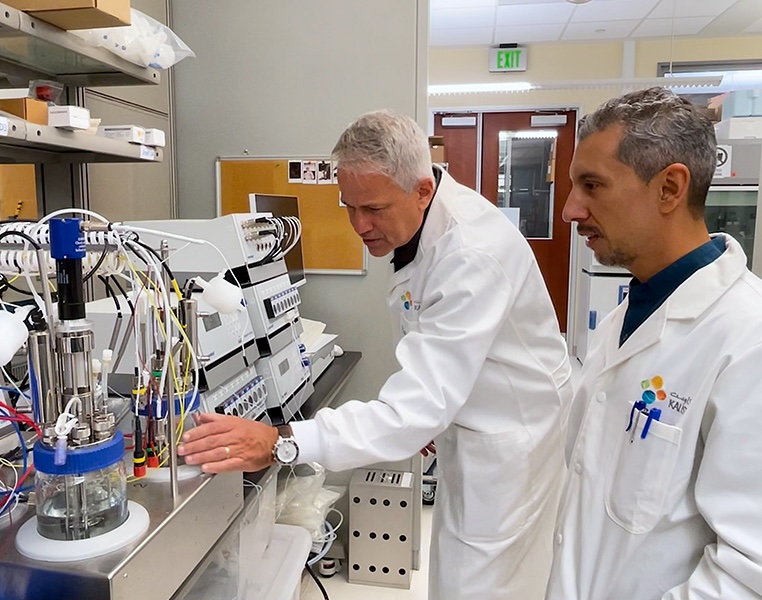

Many microbes live in diverse symbiotic communities and rely on interactions between species to survive. These communities are often disrupted by isolation techniques, causing scientists to lose the more sensitive, rare species — or miss them entirely.
“In many cases, we are simply not recreating the right ecological context,” says Rosado. “The limitation is not the methods themselves, but the fact that they were never designed to capture the full diversity we now know exists.”
Genome-based cultivation strategies have improved knowledge of the nutritional requirements needed to cultivate specific prokaryotic taxa. However, many prokaryotic genomes contain genes with unknown or poorly annotated functions, making it more difficult to understand the microbes’ physiology and predict their metabolic needs.
“Our new approach tries to reduce these challenges by using genomic information to guide cultivation, moving away from trial and error towards more informed hypotheses,” says Jiménez. “What we propose is not a new tool, but a way to connect existing approaches into a more systematic and predictive framework.
Harnessing technological advances
Analyzing and understanding thousands of microbial species in their natural environments requires a combination of multidisciplinary techniques. In their latest paper, the research team proposes combining direct environmental DNA analyses with genome-based metabolic modeling, physiological inference, and the design of tailored growth media to improve the targeted cultivation and isolation of novel taxa.
The team first proposes targeted perturbations in natural ecosystems to increase the abundance of rare or hidden microorganisms that may otherwise remain undetected. Once these microbes have been identified and collected, their genomes can be sequenced, reconstructed, and annotated. Scientists can then infer each microbe’s metabolism and potential growth requirements. Finally, this information can be used, together with AI and machine learning, to design better cultivation strategies.
However, even with a robust, reproducible framework in place, significant challenges remain, say Jiménez and Rosado.
“A large fraction of genes in microbial genomes have unknown functions, which limits how accurately we can predict physiology. While AI and machine learning are helpful, especially for gene annotation and metabolic prediction, they are not shortcuts. They depend on good data and still require experimental validation.”
Wide-reaching potential applications
This study brings together microbiology, bioinformatics, and expertise in AI and machine learning at KAUST, and represents an important step toward improving microbial cultivation.
“Once you bring new microbes into culture, you can understand their biology and begin to use it,” says Rosado. “This can translate into environmental biotechnology, agriculture, and medical applications. For example, microbes that degrade pollutants or plastics, or that support plant growth under extreme conditions, are particularly relevant in regions like Saudi Arabia.”
“We have already identified previously uncharacterized microbial taxa, discovered novel enzymes, and generated new insights into environmental microbiomes from Saudi Arabia and beyond,” says Jiménez.
“Building on these advances, the opportunity ahead is to translate this knowledge into scalable solutions,” concludes Rosado. “From human health and desert agriculture to mangrove restoration, Red Sea ecosystems, and space biology, microbiome-based approaches could support sustainable bio-based approaches aligned with KAUST priorities in energy, water, food, and health, as well as Saudi Arabia’s Vision 2030.”
Reference
- Jiménez, D.J., Marasco, R., Schultz, J., Rodriguez, C.A.D., Nogales, J., Rodriguez-R, L.M., Overmann, J. & Rosado, A.S. Discovery and cultivation of prokaryotic taxa in the age of metagenomics and artificial intelligence. The ISME Journal 20, 1 (2026). | article
You might also like

Environmental Science and Engineering
Climate action needs a tree change

Chemical Engineering
Natural solvents enable fully bio-based membranes

Environmental Science and Engineering
Plant diversity reduces the impacts of grazing pressure in drylands
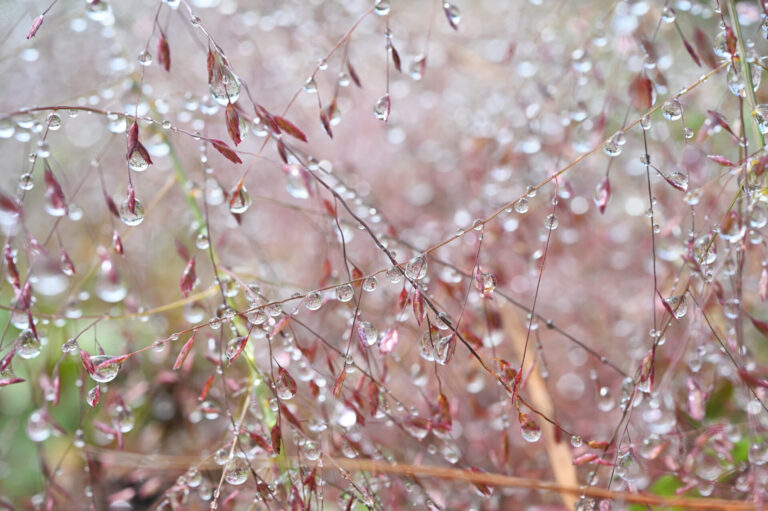

Environmental Science and Engineering
Water-repelling surfaces reveal surprising charging effects
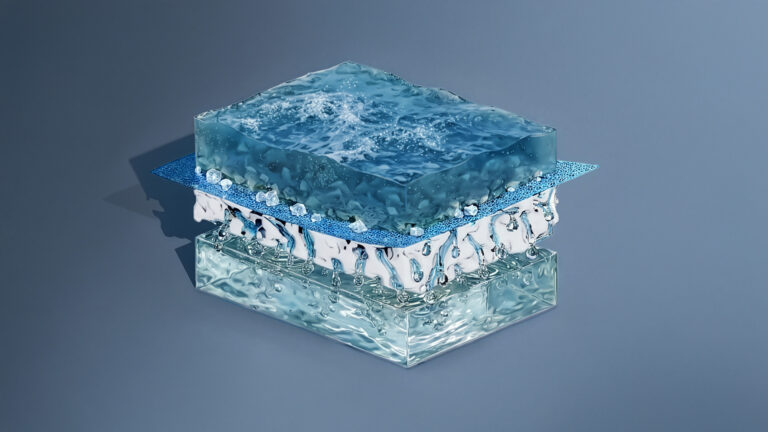

Environmental Science and Engineering
Ultrathin water repellent membrane advances desalination
Environmental Science and Engineering
Bacteria reveal hidden powers of electricity transfer
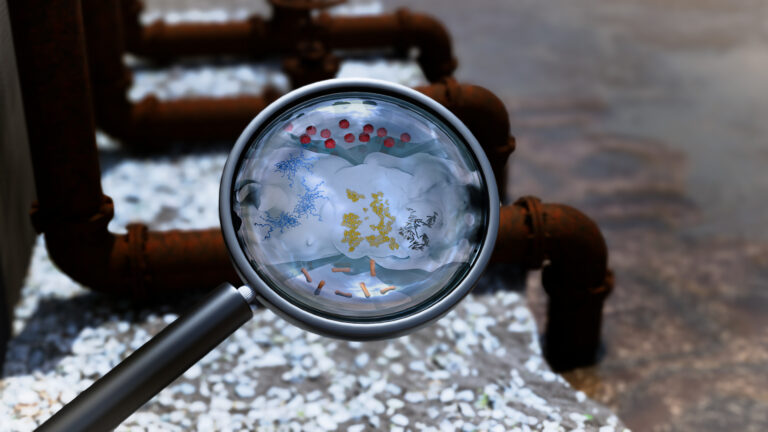

Environmental Science and Engineering
Wastewater surveillance tracks spread of antibiotic resistance
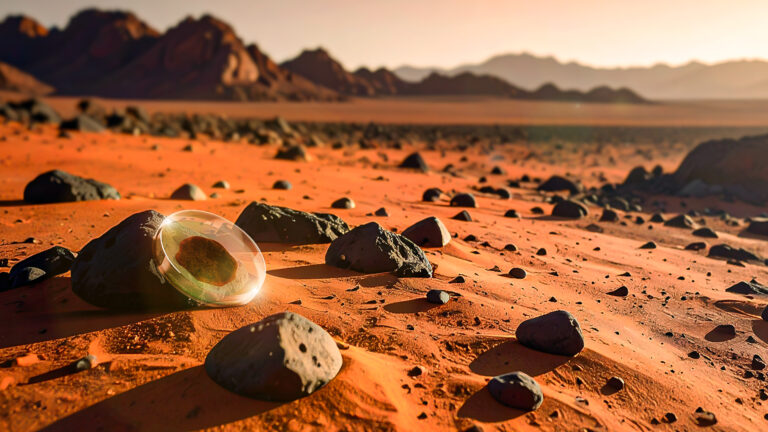

Bioscience




