Bioscience
Quenching scientific curiosity with single-molecule imaging
New experimental insights allow researchers to probe protein-DNA interactions with greater precision.
A single-molecule imaging technique, called protein-induced fluorescence enhancement (PIFE), has gained traction in recent years as a popular tool for observing DNA–protein interactions with nanometer precision. Yet, according to a new KAUST study, research laboratories have not been using the technique to its fullest potential.
KAUST graduate students work on protein-DNA analyses.
© KAUST 2019
The PIFE assay is predicated on the idea that DNA tagged with a fluorescent dye will glow brighter when proteins are bound in the vicinity. In many instances, this is true—which has led many scientists to adopt PIFE over other more labor-intensive techniques that rely on dual labeling of proteins and DNA.
But Samir Hamdan’s graduate students Fahad Rashid, Manal Zaher and Vlad-Stefan Raducanu realized that protein binding to DNA-dye complexes could sometimes have the opposite effect as well. Instead of enhancing the fluorescent signal, protein interactions can sometimes dampen the glow, depending on certain properties of the system.
Hamdan credits the curiosity of his students for making this observation and detailing how it works. Inspiration from Rashid’s previous work led the team to the phenomenon they call protein-induced fluorescence quenching (PIFQ). And as Rashid explains, “We set out to better define the conditions that lead to fluorescent booms or busts.”

Vlad-Stefan Raducanu illustrates the fluorescent (left) and quenched states of the DNA-dye complex.
© 2019 KAUST
Through a combination of experimental and computational analyses, the KAUST team showed that the initial fluorescence state of the DNA-dye complex determines whether PIFE or PIFQ will result after protein binding. Without this knowledge, the likelihood of either event becomes equivalent to a coin toss, which can jeopardize the mechanistic interpretation of laboratory results.
“When insight into this initial state is gleaned from fluorescence and structural work, the anticipation of either effect becomes experimentally feasible,” Raducanu explains.
Factors such as DNA sequence and dye position could tip the balance toward PIFE or PIFQ; the KAUST team got so good at interpreting the molecular code that they could accurately predict which would happen simply by measuring how these parameters influence the initial fluorescence state of the DNA-dye system.
“We turned every measurement into a game,” Zaher says, “And we are happy to say that our hypothesis predicted the outcome more than 90 percent of the time!”
These novel insights should dramatically expand the reach and experimental promise of this powerful single-molecule imaging tool, predicts Raducanu. “By introducing PIFQ, we offer researchers in the field the possibility to address several biological questions where PIFE might not have been witnessed,” he says.
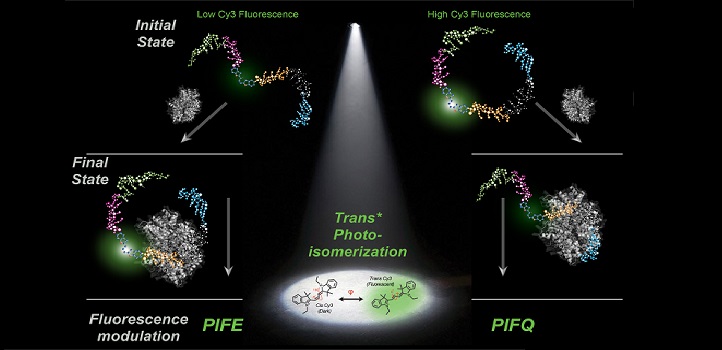
The initial state of the DNA-dye complex determines whether protein binding enhances or quenches the fluorescence signal.
© 2019 KAUST
Scientists may also opt to combine PIFE and PIFQ to decipher multistep and multiprotein processes with just a single DNA-dye construct.
“Taking into consideration the context-dependent nature of fluorescence modulation in the DNA-dye system opens the door to many possibilities in experimental design that could be tailored to researchers’ needs,” Zaher says.
“We now anticipate that interpretation of data and attribution of molecular events from single-molecule data will become easier and more precise,” Rashid adds.
References
- Rashid, F., Raducanu, V.S., Zaher, M.S., Tehseen, M., Habuchi, S. & Hamdan, S.M. Initial state of DNA-Dye complex sets the stage for protein induced fluorescence modulation. Nature Communications 10, 2104 (2019).| article
You might also like

Bioscience
Single small-molecule model reveals insights into human embryo development

Bioscience
Can AI finally bring order to biology’s data deluge?

Bioengineering
Bio-inspired network structures for next-generation AI

Bioscience
Robust workflow built for chemical genomic screening

Bioscience
Cell atlas offers clues to how childhood leukemia takes hold

Bioscience
Hidden flexibility in plant communication revealed

Bioscience
Harnessing the unintended epigenetic side effects of genome editing

Bioscience




