Bioscience
DNA division can slow to a halt
Termination sites of DNA are shown to stop slow-moving replication forks but not faster ones.
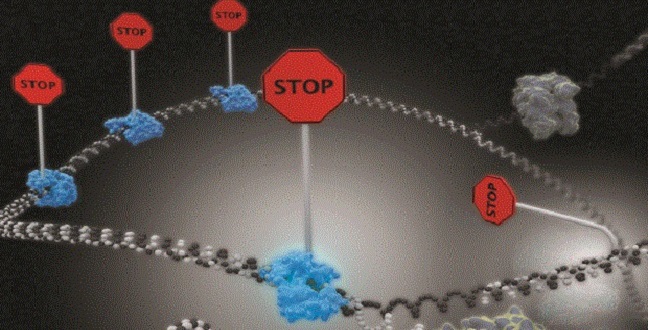
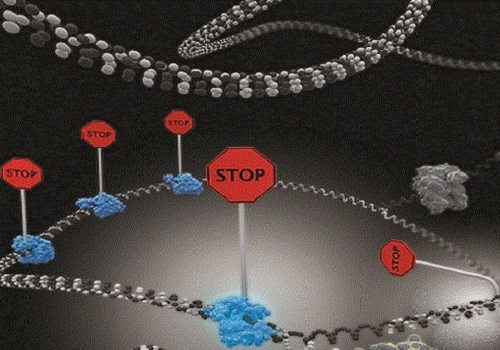
Single-molecule imaging reveals that DNA replication termination in E. coli is mediated by kinetic competition between speed of strand separation by the replisomal helicase and rearrangement of Tus−Ter interactions during separation of the first six base pairs of Ter. Multiple termination sites are required to insure fork stoppage.
© 2015 KAUST
A key mystery of the DNA replication process has been unraveled by researchers from KAUST.
Before a bacterium can divide, it must make a copy of its genetic material, the circular DNA molecules that resemble bunched rubber bands, through a process called DNA replication. In this process, the two strands of DNA making up the circular DNA molecule unwind and separate to become templates for generating new strands.
To ensure the process is well regulated, the bacterium has set a number of ‘roadblocks’, or termination sites on the DNA, to ensure the permanent stoppage of replication forks, Y-shaped structures formed between the strands as the DNA molecule splits.
The Nature study, led by KAUST Ph.D. student Mohamed Elshenawy and Associate Professor Samir Hamdan from KAUST’s Division of Biological and Environmental Science and Engineering, along with colleagues from the University of Wollongong in Australia, showed why termination sites were able to permanently stop replication forks in vitro, while in living bacteria, more than 50 percent of the replication forks that moved towards the termination site continued synthesis without stopping1.
A termination site comprises a 23-base pair termination sequence (Ter) bound to the protein terminus utilization substance (Tus). Tus−Ter is unusual in that it acts like the ratcheting knot on a climbing rope by allowing progression of replication forks from one direction but not the other. This polarity sets up a ‘trap’ that allows the first arriving fork to enter but not to leave the terminus region until the other fork has arrived.
Hamdan and his team suspected that the energy from movement (or kinetics) might be acting on these termination sites. They used single-molecule imaging to record molecular movies that zoomed in with high temporal and spatial resolution on the fate of Escherichia coli replication forks as they approached a termination site from either direction.
Their results showed that efficiency of fork arrest is weakened by kinetic competition between the rate of strand separation by the helicase motor at the fork and the rate of rearrangement of Tus−Ter interactions that maintain Tus’s strong grip on the DNA. This means that faster moving forks beat Tus−Ter rearrangement and displace Tus, while slower ones are effectively blocked.
This resolves a long-standing mystery that has clouded our understanding of DNA replication, and also has important implications for all domains of life.
“These findings are striking for the field of enzymology,” Hamdan stated. He noted that the demonstration that the rates of individual enzymes fluctuate during catalysis and that rates can differ among presumably identical enzyme molecules are both novel contributions of single-molecule imaging to biology.
“This study demonstrated for the first time that these intrinsic properties of enzyme molecules actually impact biology,” explained Hamdan.
The communication between molecular motors and double-stranded DNA binding proteins is a common feature in DNA replication, repair, recombination and transcription and also in instances where conflict occurs between these processes. The evolution of different responses to the average rate of different molecular motors could regulate the communication among these processes, Hamdan said.
References
-
Elshenawy, M.M. Jergic, S., Xu, Z-Q., Sobhy, M.A., Takahashi, M., Oakley, A.J., Dixon, N.E. & Hamdan, S.M. Replisome speed determines the efficiency of the Tus−Ter replication termination barrier. Nature 524, 394-398 (2015).| article
You might also like

Bioengineering
Bio-inspired network structures for next-generation AI

Bioscience
Robust workflow built for chemical genomic screening

Bioscience
Cell atlas offers clues to how childhood leukemia takes hold

Bioscience
Hidden flexibility in plant communication revealed

Bioscience
Harnessing the unintended epigenetic side effects of genome editing

Bioscience
Mica enables simpler, sharper, and deeper single-particle tracking

Bioengineering
Cancer’s hidden sugar code opens diagnostic opportunities
Bioscience



