Computer Science
Peak performance for proteins
An improved peak fitting procedure enables a better determination of protein structures.
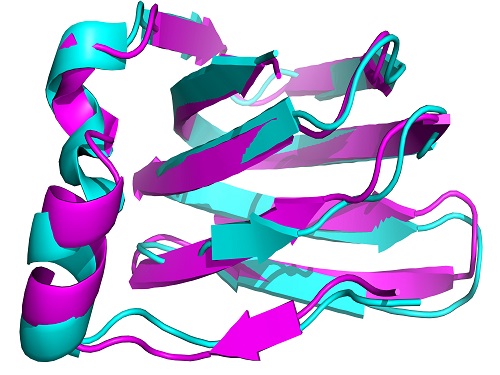


An automatically calculated structure for the protein TM1112, derived from NMR spectra.
© 2015 KAUST
A method to more accurately determine the structure of proteins using nuclear magnetic resonance (NMR) spectroscopy has been developed by researchers at KAUST.
Proteins play a key role in cell function and vital to a protein’s performance is its molecular structure, which usually assumes a complex three-dimensional shape (see image).
A common method of identifying protein structures uses x-ray scattering techniques: but this approach is imperfect because it requires proteins to be available in crystalline form. NMR is an improvement because it can resolve molecular structures in cells, and is therefore uniquely suited to also measure the dynamics of proteins.
NMR measures an atom’s energetic states with high sensitivity and enables researchers to study the bonds between atoms in a molecule and even the local atomic geometry around the atoms. A protein can consist of many thousands of atoms that contribute to NMR signals, and so the measured spectra are extremely complex. Interpreting the protein structure from these is difficult and requires sophisticated computer models.
An obstacle to analyzing NMR spectra is that some of the signals are very small, and detecting them against a noisy background is difficult. There are several established computer algorithms for such peak detection. In these, the NMR signal is divided into small sections, known as ‘windows’. Then the algorithm determines the background for each window and extracts the NMR peaks.
The difficulty is to find the right tradeoff, explains research leader Xin Gao, from the Computer, Electrical and Mathematical Science and Engineering Division at KAUST.
“On one hand, you need to remove the noise from the spectrum as much as possible,” says Gao. “On the other hand, there is the risk of accidentally removing real signals.”
The filtering algorithm developed by the researchers is an extension of prior schemes and makes it easier to find the right balance. It features a better detection of the edges of peaks in the spectra, which other techniques tend to smooth over, thus making signals more difficult to isolate. Moreover, the new algorithm is also independent of the window size used, which represents a considerable practical advance.
The new detection leads to a better real-time study of protein dynamics, comments Gao. “We already have an in-house automated system that can determine small and medium size protein structures without human intervention. In future, we hope that by combining the proposed noise removal filter, and our in-house tools, we will be able to push the limit to larger proteins.”
References
- Cannistraci, C.V., Abbas, A. & Gao, X. Median Modified Wiener Filter for nonlinear adaptive spatial denoising of protein NMR multidimensional spectra. Scientific Reports 5, 8017 (2015).| article
You might also like

Bioscience
Can AI finally bring order to biology’s data deluge?

Computer Science
AI-based hydrogen plant models improve power grid stability

Bioengineering
Bio-inspired network structures for next-generation AI

Computer Science
Green quantum computing takes to the skies

Computer Science
Probing the internet’s hidden middleboxes
Bioscience
AI speeds up human embryo model research

Computer Science
Improving chip design on every level

Computer Science



