Plant Science
Tending the future of data analysis with MVApp
New app aims to improve the statistical analysis of large datasets in plant science and beyond.

The vast datasets generated by modern plant-science technologies require clever data-mining methods to extract useful information. Now, KAUST researchers have developed MVApp—an open-source, online statistics platform for conducting multivariate analyses of these intricate data.
The recent development of high-throughput phenotyping techniques has rapidly produced huge datasets on the characteristics of plants. These multivariate data hold crucial details about plant physiology: how a plant responds to different environments and how a plant’s growth patterns and potential yields change—all of which are valuable in developing sustainable agriculture and ensuring food security.
“Our experiments typically include measurements of thousands of plants every day for multiple traits, from leaf size through to salt-stress tolerance or resistance to plant pathogens,” says Magdalena Julkowska, a research scientist working in Mark Tester’s lab at KAUST. “These data are extremely powerful, but overwhelming to sieve through, she adds, “As a team, we know the struggles that come with large data analyses, and we figured if we struggle, then others must too.”

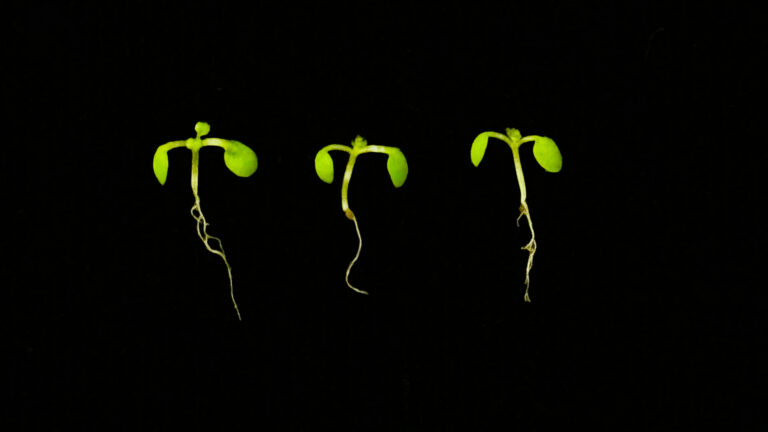
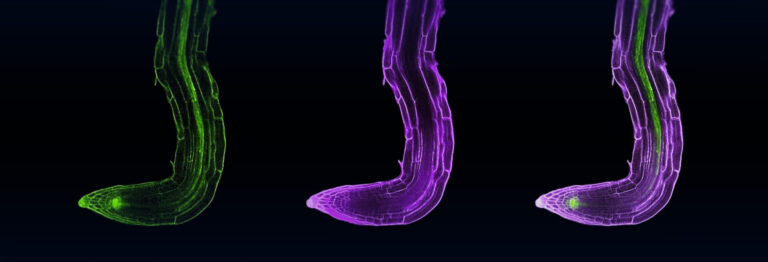
The team typically uses small flowering plants called Arabidobsis thaliana as a model plant for their research.
© 2019 KAUST
The team built MVApp using R-language—a popular tool for statistical analyses—and incorporated the most pertinent R packages for analyzing phenotyping data. MVApp can be used with datasets of different sizes, from exploratory analyses of large-scale natural diversity to smaller-scale projects comparing mutant phenotypes to wild-type plants.
The team also incorporated a technique called quantile regression into MVApp—this specialist tool is used in other fields but has not yet reached its full potential in plant science.
“When we screen populations of hundreds of diverse plant accessions, originating from different parts of the globe, the plants that yield well might have different traits contributing to yield than the plants that produce low yields,” says Julkowska. “Let’s say you’d like to explain the yield of a specific plant type by its biomass and water use—quantile regression can help quantify how much each trait is contributing to your main trait of interest.”
MVApp produces comprehensive, easy-to-follow outputs, and generates attractive, publication-ready figures that have clear links back to the raw data used to create them. The MVApp team are passionate about improving data transparency and streamlining data curation, ensuring that every scientist can produce valuable, reproducible results.
MVApp can be used with datasets of different sizes: from exploratory analyses of large-scale natural diversity to smaller-scale projects comparing mutant phenotypes to wild-type plants. Reproduced with permission from reference 1 © 2019 American Society of Plant Physiologists
“We hope that MVApp will help the entire scientific community—not just plant scientists—to become familiar with various statistical methods and to know how to implement them, particularly with big data,” says Julkowska. “We welcome feedback and hope users will help us improve MVApp.”
References
- Julkowska, M.M., Saade, S., Agarwal, G., Gao, G., Pailles, Y. Morton, M.J.L., Awlia, M.& Tester, M. MVApp – Multivariate analysis application for streamlined data analysis and curation. Plant Physiology 00235 (2019).| article
You might also like

Plant Science
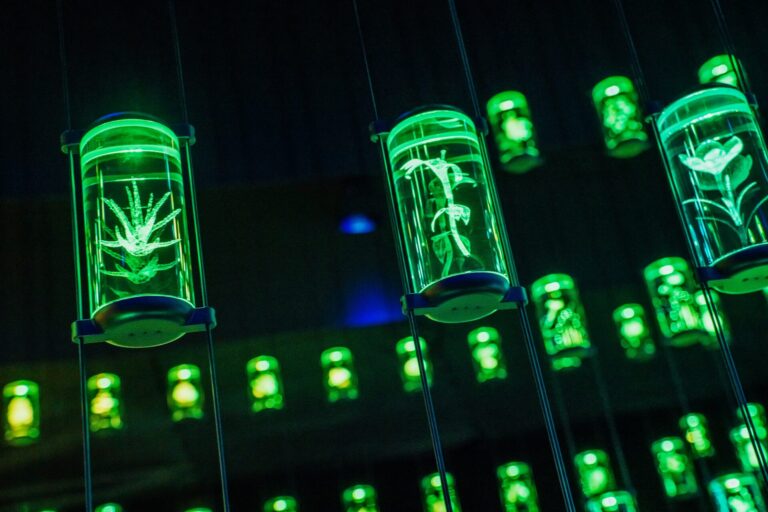
Tiny DNA fragments, big agricultural insights

Plant Science
Coevolution of the microbiome with its host plant supports environmental adaptation

Bioscience
Hidden flexibility in plant communication revealed

Plant Science
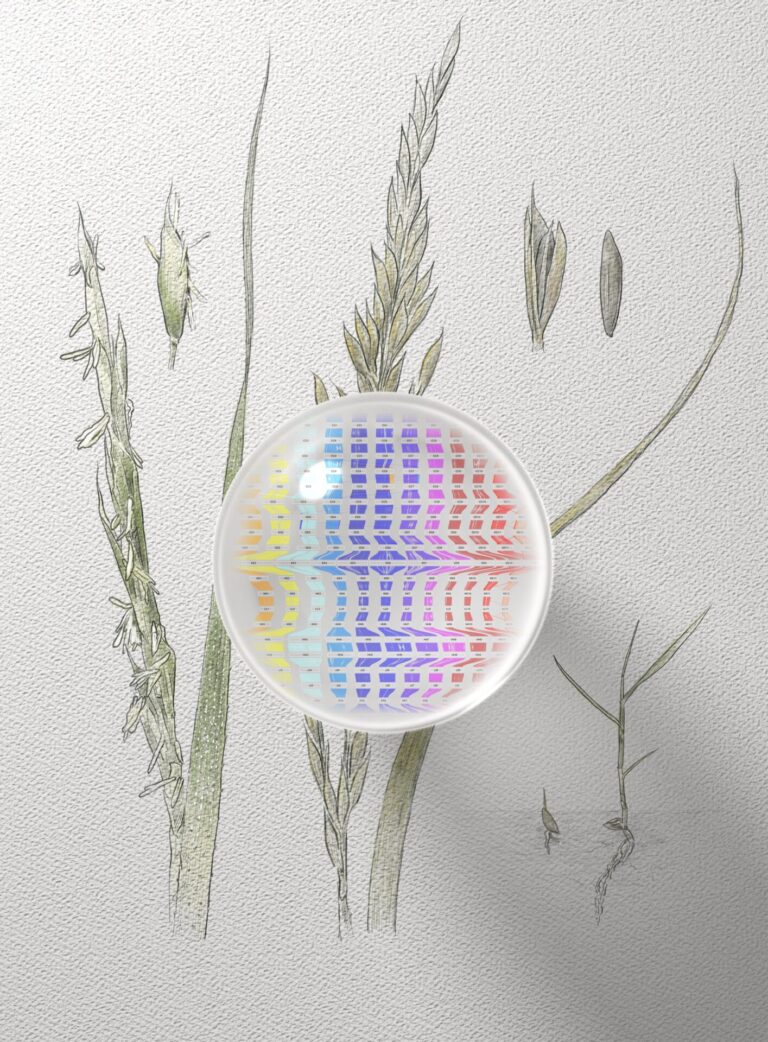
Reference genomes for rice’s wild relatives may boost future crops
Bioscience
Digging into the world of plant-growth-promoting microbes
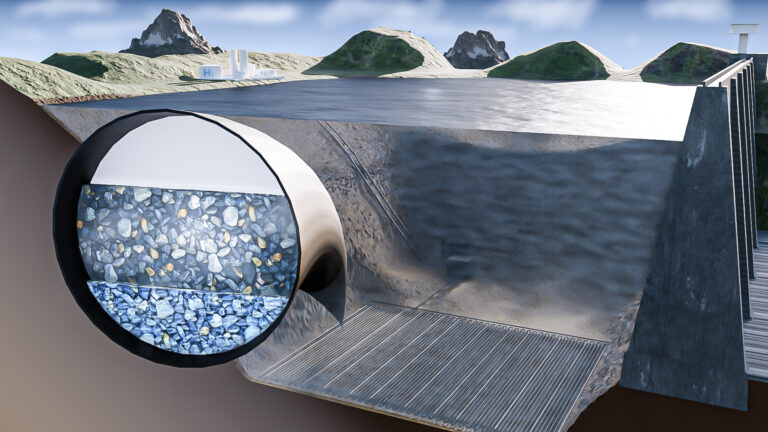

Environmental Science and Engineering
Hydrogen storage solution could lie in lakes

Bioscience
Unraveling modern bread wheat from the genes up

Bioscience




