Applied Mathematics and Computational Sciences
Mining Red Sea bacteria for industrial potential
Genome sequencing of two Red Sea bacteria highlights their potential as industrial workhorses.
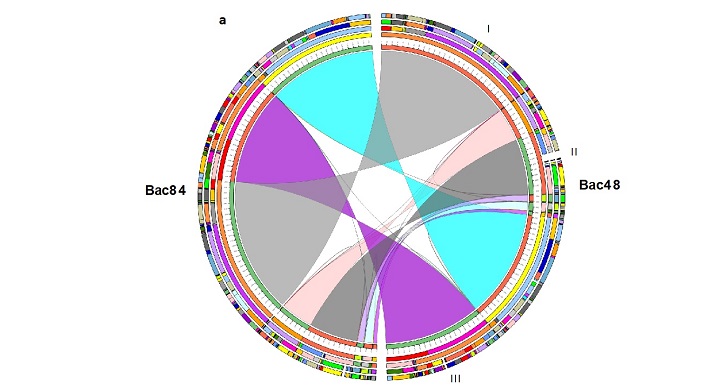
Analyses of two bacterial strains from the Red Sea show they are enriched with gene clusters with potential to activate the synthesis of a wide range of industrially useful compounds, from novel antibiotics, anticancer agents and pigments to those useful for crop protection and the food industry.
Bacteria are a rich resource for bioactive chemical compounds and Magbubah Essack, of KAUST’s Computational Bioscience Research Center, explains that bacterial strains able to withstand the Red Sea’s highly saline, warm waters were anticipated to produce sturdy enzymes suited for industrial applications.
The team sequenced the genomes of two Bacillus species: B. paralicheniformis Bac48 collected from mangrove mud and B. paralicheniformis Bac84 collected from a microbial mat in the Rabigh Harbor Lagoon on Saudi Arabia’s west coast. These two were compared with the documented genomes of three other B. paralicheniformis and nine B. licheniformis strains. The Red Sea strains had a higher number of gene clusters associated with bioactive compound synthesis compared to the other Bacillus strains.

The research team report the complete circular and annotated genomes of two Red Sea strains, B. paralicheniformis Bac48 isolated from mangrove mud and B. paralicheniformis Bac84 isolated from microbial mat collected from Rabigh Harbor Lagoon in Saudi Arabia. (Front from l-r: Takashi Gojobori, Vladimir Bajic, Heribert Hirt; Back from l-r: Magbubah Essack, Ameerah Bokhari, Salim Bougouffa, Xin Gao, Stefan Arold.)
© 2018 KAUST
The team also report the first use of a computer program to identify a gene cluster in strains of the B. paralicheniformis species, in this case B. paralicheniformis Bac48, called trans-acyltransferase nonribosomal peptide synthetase/polyketide synthase, which is associated with the production of a specific group of compounds.
“The findings affirm the premise that the Red Sea is a metabolically unique environment worthy of exploration for efficient microbes that can be used as biotechnological hosts,” says computational bioscientist Ghofran Othoum, the first author of the study. “Also, our computational exploratory approach showed the strength of computer modeling methods in applications that require ranking biological systems for biotechnological use.”
Future research will focus on seeking out which bioactive compounds are produced by the two Red Sea strains.
References
- Othoum, G., Bougouffa, S., Razali, R., Bokhari, A., Alamoudi, S., Autunes, A., Xao., G., Hoehndorf, R., Arold, S.T., Gojobori, T., Hirt, H., Mijakovic, I., Bajic, V.B. Lafi., F.F. & Essack, M. In silico exploration of Red Sea Bacillus genomes for natural product biosynthetic gene clusters. BMC Genomics 19, 382 (2018).| article
You might also like

Applied Mathematics and Computational Sciences
Resilient renewable energy networks designed for the desert
Applied Mathematics and Computational Sciences
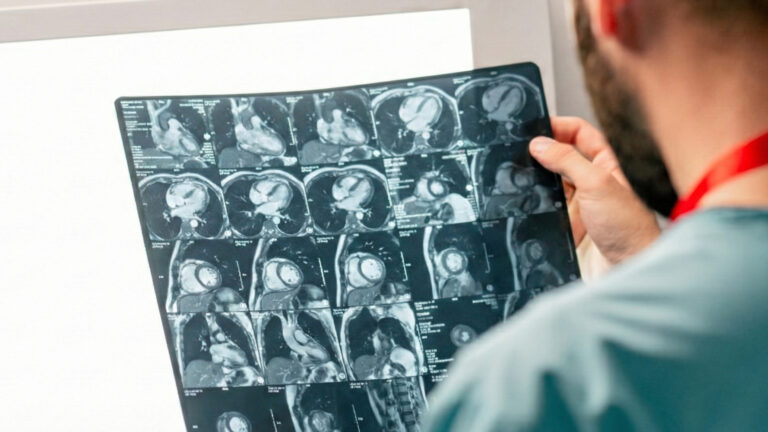
Smarter MRI image analysis for the whole heart

Applied Mathematics and Computational Sciences
Realistic scenario planning for solar power
Applied Mathematics and Computational Sciences
Bringing an old proof to modern problems

Applied Mathematics and Computational Sciences
Accounting for extreme weather to boost energy system reliability
Applied Mathematics and Computational Sciences
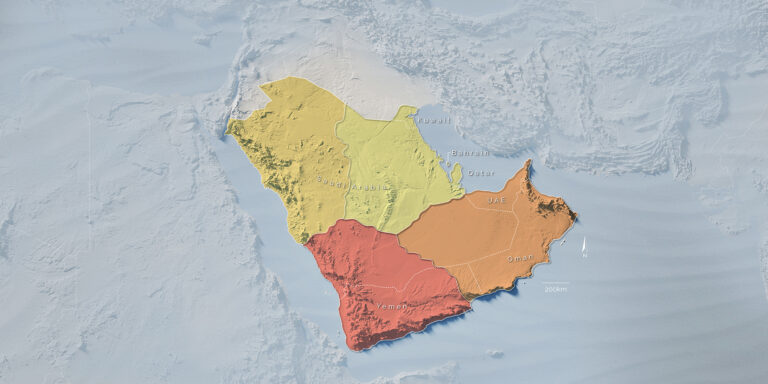
Past and future drought patterns across the Arabian Peninsula

Applied Mathematics and Computational Sciences
New pattern for underwater resonators

Applied Mathematics and Computational Sciences




