Plant Science
Future for crop breeding is CRISPR and clear
Precision molecular tools derived from bacterial defence systems are transforming the theory and practice of genetic modification.

Plant scientists, breeders and growers alike are always seeking new weapons in the fight against disease. One of the most exciting tools in their armoury is the CRISPR/Cas system, the subject of a pioneering research program in the KAUST Desert Agriculture Initiative.
CRISPR/Cas technology can be used to remove, add, alter or deactivate specified sections of an organism’s DNA. Compared to other genome editing methods, CRISPR/Cas is considered faster, cheaper and more accurate.
The system originated naturally in bacteria, delivering adaptive immunity against viral pathogens. Bacteria infected by a virus are able to copy parts of the viral DNA into their own genome within a clustered regularly interspaced short palindromic repeat (CRISPR) array, providing them with a memory of the virus. This memory primes them to defend against future attacks. To do this, small molecules known as CRISPR RNAs (crRNAs), produced from the array, bind to the viral DNA1, which CRISPR-associated (Cas) proteins, attracted by the crRNA, then attack and destroy. The mechanism has already been lauded as a promising weapon against diseases, such as HIV/AIDS.
In 2015, researchers led by postdoc Zahir Ali, under Magdy Mahfouz, successfully implemented CRISPR technology with the Cas9 protein in tobacco plants. They showed that Cas9 could deactivate not just one, but multiple viruses simultaneously if plants were primed with multiple crRNAs, each complementing a different virus2. This ground-breaking study highlighted the potential of the CRISPR/Cas system for crop protection.
The CRISPR/Cas9 system is, says Mahfouz, “one of the few that has made it into the lab.” However, as more microbial genomes have been sequenced, many tens of Cas proteins have been identified. The KAUST team is also addressing the more challenging question of defence against RNA-based viruses. Although the genome of most organisms comprises double-stranded DNA, many viral genomes are held in a single-stranded RNA form. In fact, the majority of destructive plant viruses are RNA-based, and thus, Mahfouz points out, “There was high demand for a CRISPR-based RNA-targeting technology.”
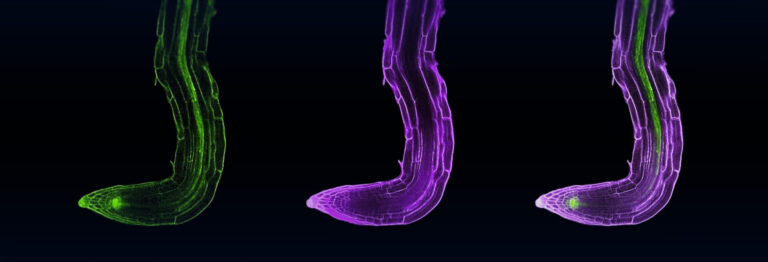
Writing in Genome Biology3, Mahfouz describes how the enzyme Cas13a, obtained from Leptotrichia shahii bacteria, cleaves RNA. Postdoc Rashid Aman says, “it has unique features not present in other known Cas proteins.” They successfully rewired CRISPR/Cas13a to provide resistance to turnip mosaic virus (TuMV) by creating tobacco plants that produce both Cas13a and four crRNAs complementary to parts of the TuMV genome. To visualise their findings, they used TuMV particles engineered to produce a fluorescent green protein. Plants expressing the Cas13a system showed around a 50 percent decrease in fluorescence, indicating that the CRISPR/Cas13a system can provide defence against RNA viruses in crop species.
CRISPR/Cas “has been transformative in the field of agriculture, revolutionising plant breeding,” says Mahfouz. But each system requires extensive optimisation in the lab. “There is still much to be done to take this technology to the next stage,” says Ahmed Mahas, a graduate student in Mahfouz’s lab. In the CRISPR/Cas13a system, different crRNAs showed significant variation in effectiveness, suggesting that refining their sequences may be vital to success. The KAUST team continue to explore the biology of the system to better understand its mechanism of action and to find Cas13 variants with stronger anti-RNA activity.
The final hurdle to widespread implementation of CRISPR/Cas technology is public and political resistance to genetically modified organisms. For this reason, says Mahfouz, “we are now focusing on building CRISPR/Cas technologies without incorporating any foreign DNA to produce enhanced plants that are not considered genetically modified.” Also, says Mahas, “We are now expanding our work on CRISPR/Cas13 for a variety of other applications.” The Cas13a system has important properties beyond RNA cleaving: it appears to play a role in processing the crRNA molecules that direct it to its target sequences. This may be important for engineering crops that are able to resist multiple RNA- or DNA-based viruses at the same time. In addition, some Cas13 variants show collateral activity and could potentially act as a broad-spectrum RNA-destroying machine, which could limit pathogen spread by triggering the death of infected cells. The CRISPR/Cas system may also be used as a diagnostic tool to identify pathogens, and the researchers in the Mahfouz lab are seeking to identify the Cas13 variants that might be best-suited to this role4.
Together, CRISPR/Cas9 and CRISPR/Cas13a provide complementary tools for editing DNA and its crucial partner—RNA. “Almost all biological processes involve RNA,” Mahfouz says, “CRISPR/Cas13 holds great promise as a powerful tool for RNA manipulation and RNA virus targeting in plants with potential to mitigate some of the major agricultural challenges around the world.”

Tobacco plants producing Cas13a are protected against turnip mosaic virus (TuMV, green florescence). Upper left shows infected leaves with no crRNA and upper right shows infected leaves with a control crRNA that does not bind TuMV. The remaining four images show varying reductions of fluorescence provided by four different TuMV-specific crRNAs.
© 2018 Magdy Mahfouz
References
- Ali, Z., Mahas, A., & Mahfouz, M. CRISPR/Cas13 as a tool for RNA interference. Trends in Plant Science 23, 374—378 (2018).| article
- Ali, Z., Abulfaraj, A., Idris, A., Ai, S., Tashkandi, M. et al. CRISPR/Cas9-mediated viral intereference in plants. Genome Biology 16, 238 (2015).| article
- Aman, R., Ali, Z., Butt, H., Mahas, A., Aljedaani, F. et al. RNA virus interference via CRISPR/Cas13a system in plants. Genome Biology 19, 1 (2018).| article
- Mahas, A., Neal Stewart, C., Jr. & Mahfouz, M.M. Harnessing CRISPR/Cas systems for programmable transcriptional and post-transcriptional regulation. Biotechnology Advances 36, 295—310 (2017).| article
You might also like

Plant Science
Coevolution of the microbiome with its host plant supports environmental adaptation

Bioscience
Hidden flexibility in plant communication revealed

Plant Science
Reference genomes for rice’s wild relatives may boost future crops
Bioscience
Digging into the world of plant-growth-promoting microbes

Environmental Science and Engineering
Hydrogen storage solution could lie in lakes

Bioscience
Unraveling modern bread wheat from the genes up

Bioscience
Why do some plants thrive in saline conditions?
Bioengineering




